A Hidden Markov Model (HMM) extends the mixture model by allowing the latent state to evolve over time according to a Markov chain. Where a mixture model treats each observation independently, an HMM captures temporal structure: the hidden state at time depends on the state at time .
This chapter covers:
The HMM generative model
Exact inference via the forward-backward algorithm
Learning parameters with EM (the Baum-Welch algorithm)
Source
import torch
import torch.distributions as dist
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
palette = list(plt.cm.Set2.colors)
torch.manual_seed(305)
<torch._C.Generator at 0x7fe728fdfa30>From Mixture Models to HMMs¶
Recall the Gaussian mixture model from Chapter 8:
The i.i.d. assumption means each data point is assigned to a cluster independently of its neighbours. For time-series data this is often unrealistic: a speaker stays in the same phoneme for multiple frames; a mouse stays in the same behavioural state for seconds.
The HMM replaces the i.i.d. prior on with a Markov chain:
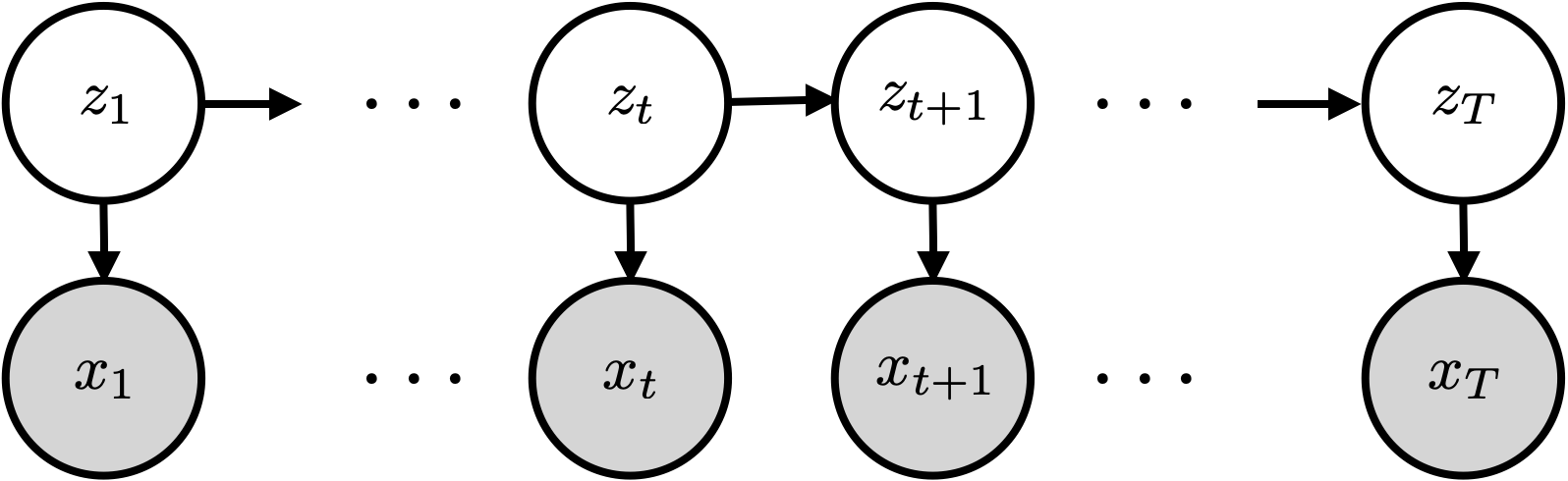
Graphical model for an HMM. The hidden states follow a Markov chain; each observation depends only on the current state .
The joint distribution factors as:
We call this an HMM because the hidden states follow a Markov chain, .
The Three Components¶
An HMM is defined by three ingredients:
Initial distribution — which state to start in.
Transition matrix (row-stochastic) — .
Emission distribution — Gaussian, categorical, Poisson, etc.
Example: The Dishonest Casino¶
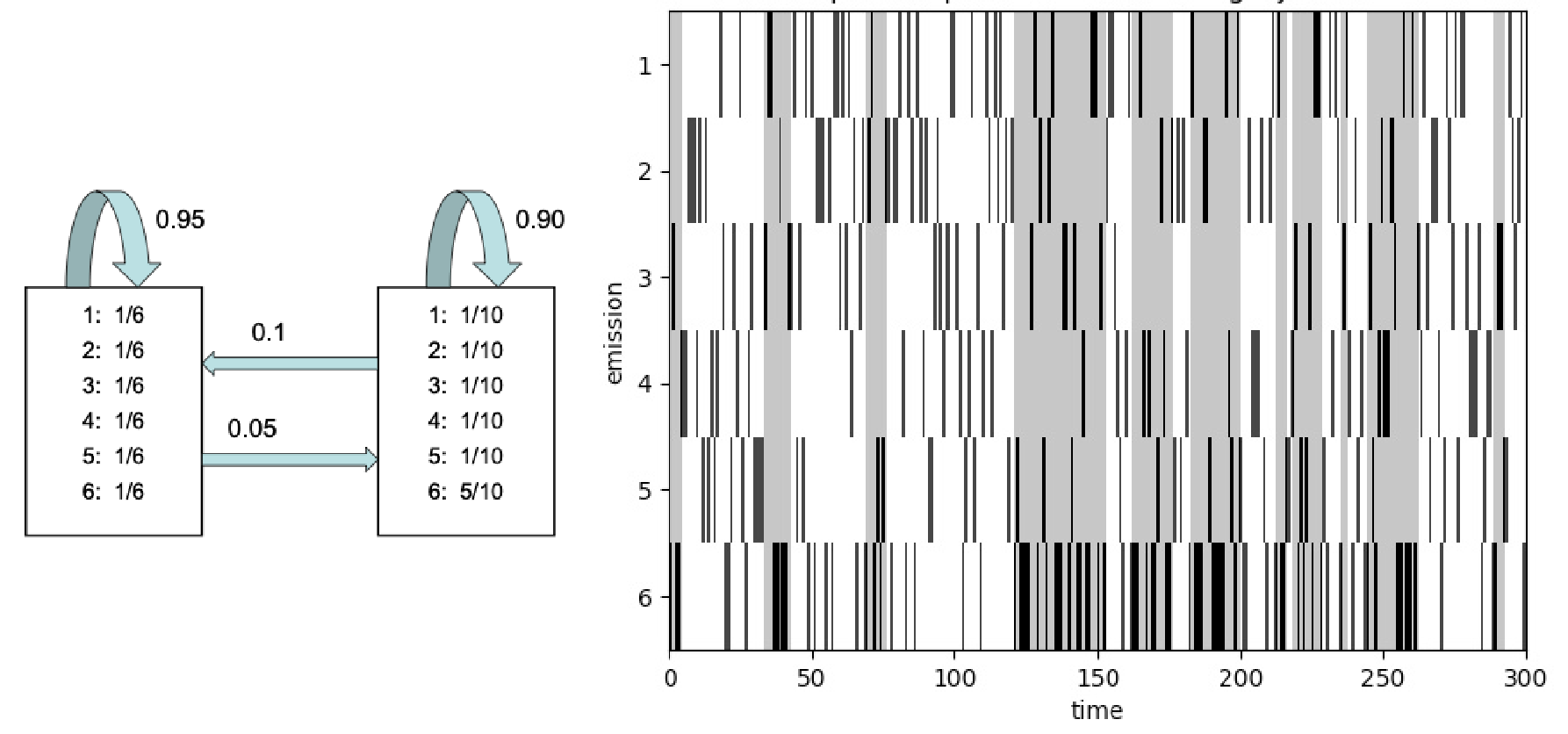
An occasionally dishonest casino that switches between a fair die () and a loaded die (, ). Figure from dynamax.
Example: Splice Site Recognition¶
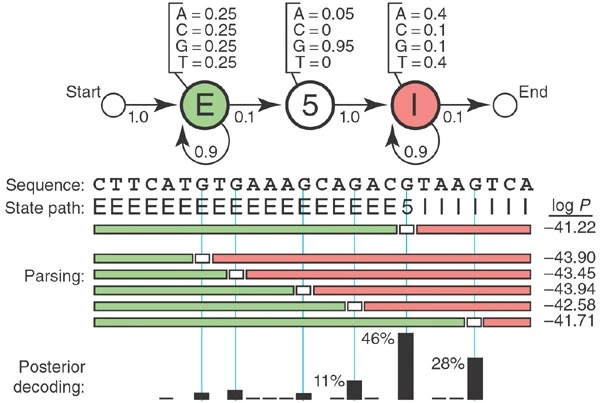
A toy HMM for parsing a genome to find 5'' splice sites. Figure from Eddy, 2004.
Example: Behavioural Segmentation of Video¶
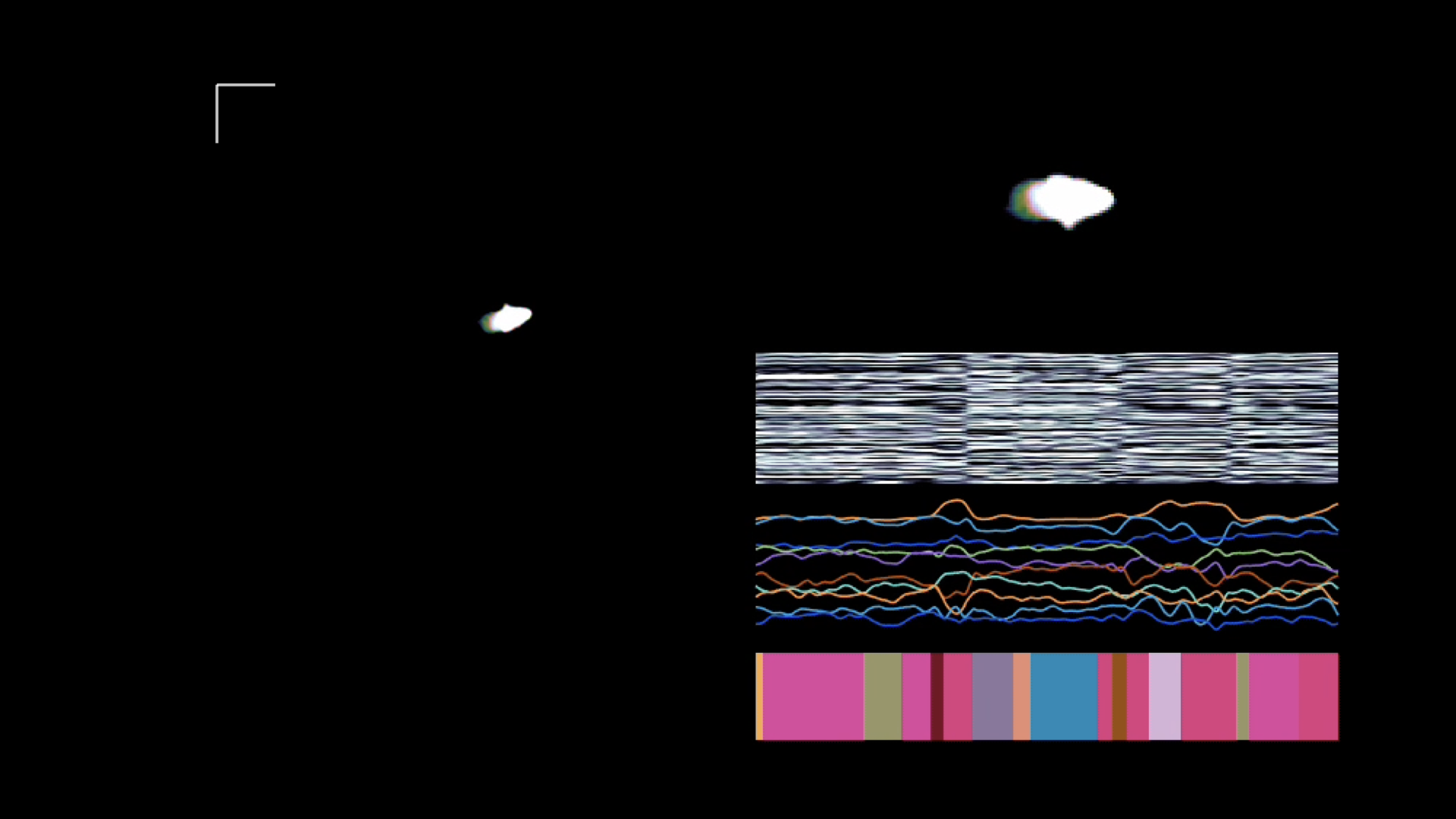
Segmenting videos of freely moving mice with an autoregressive HMM. Figure from Wiltschko et al., 2015.
Inference Goals¶
Given observations and parameters , we want:
| Quantity | Name |
|---|---|
| Filtering — current state given past | |
| Smoothing — current state given all data | |
| Pairwise smoothing — needed for EM | |
| Viterbi — most likely path | |
| Marginal likelihood — needed for learning |
Why is naive enumeration expensive? Marginalising requires summing over state sequences. Message passing reduces this to by processing one time step at a time, exploiting the Markov structure.
The Forward Algorithm¶
Define the forward message at time as the joint probability of being in state and observing :
Base case: .
Recursion:
Let . In matrix form:
Normalizing for numerical stability. Re-normalize after each step:
The normalizers have a clean interpretation: is the predictive distribution , and . Therefore:
The Backward Algorithm and Smoothing¶
Define the backward message as the probability of future observations given the current state:
Base case: for all .
Recursion:
or in matrix form: .
Posterior Marginals (Smoothing)¶
Together the forward and backward passes give the forward-backward algorithm Rabiner & Juang, 1986.
Posterior Pairwise Marginals¶
These are needed for the EM M-step below.
EM for HMMs (Baum-Welch)¶
The ELBO for an HMM decomposes over the three parameter groups:
where (posterior marginals) and (posterior pairwise marginals).
E-step: Run the forward-backward algorithm to compute and .
M-step: The ELBO separates, giving closed-form updates:
For exponential family emissions, the weighted MLE for requires only the expected sufficient statistics — the same structure as the mixture model M-step.
# ── Forward-backward algorithm ────────────────────────────────────────────────
def forward_pass(log_pi0, log_P, log_likes):
'''Normalized forward messages.
Convention (matching the lecture):
log_alphas[t, k] = log p(z_t=k | x_{0:t-1}) (predictive, normalized)
log_norms[t] = log p(x_t | x_{0:t-1})
Args:
log_pi0: (K,) log initial distribution
log_P: (K, K) log_P[i,j] = log p(z_{t+1}=j | z_t=i)
log_likes: (T, K) log p(x_t | z_t=k)
Returns:
log_alphas (T, K), log_norms (T,)
'''
T, K = log_likes.shape
log_alphas = torch.zeros(T, K)
log_norms = torch.zeros(T)
log_alphas[0] = log_pi0 - torch.logsumexp(log_pi0, 0)
log_norms[0] = torch.logsumexp(log_alphas[0] + log_likes[0], 0)
for t in range(1, T):
log_joint = log_alphas[t-1] + log_likes[t-1] # (K,)
log_alpha_t = torch.logsumexp(log_joint[:, None] + log_P, 0) # (K,)
log_alphas[t] = log_alpha_t - torch.logsumexp(log_alpha_t, 0)
log_norms[t] = torch.logsumexp(log_alphas[t] + log_likes[t], 0)
return log_alphas, log_norms
def backward_pass(log_P, log_likes):
'''Normalized backward messages.
Args:
log_P: (K, K)
log_likes: (T, K)
Returns:
log_betas (T, K), with log_betas[T-1] = 0 (base case beta_T = 1)
'''
T, K = log_likes.shape
log_betas = torch.zeros(T, K) # base: beta_{T-1} = 1
for t in range(T - 2, -1, -1):
log_joint = log_likes[t+1] + log_betas[t+1] # (K,)
log_beta_t = torch.logsumexp(log_P + log_joint[None, :], 1) # (K,)
log_betas[t] = log_beta_t - torch.logsumexp(log_beta_t, 0)
return log_betas
def posterior_marginals(log_alphas, log_betas, log_likes):
'''gamma_t(k) = p(z_t=k | x_{0:T-1}). Returns (T, K).'''
log_gamma = log_alphas + log_likes + log_betas
log_gamma = log_gamma - torch.logsumexp(log_gamma, 1, keepdim=True)
return log_gamma.exp()
def posterior_pairs(log_alphas, log_betas, log_likes, log_P):
'''xi_t(i,j) = p(z_t=i, z_{t+1}=j | x_{0:T-1}). Returns (T-1, K, K).'''
T, K = log_alphas.shape
log_xi = (log_alphas[:-1, :, None]
+ log_likes[:-1, :, None]
+ log_P[None]
+ log_likes[1:, None, :]
+ log_betas[1:, None, :])
log_xi = log_xi - torch.logsumexp(log_xi.reshape(T-1, -1), 1).reshape(T-1, 1, 1)
return log_xi.exp()
# ── EM for an HMM with categorical emissions ──────────────────────────────────
def cat_log_likes(x, log_theta):
'''log p(x_t | z_t=k) for categorical emissions.
Args:
x: (T,) integer observations in {0, ..., V-1}
log_theta: (K, V) log emission probabilities
Returns:
(T, K)
'''
return log_theta[:, x].T
def em_hmm_cat(x, K, V, num_iters=60, seed=0):
'''Baum-Welch EM for a categorical-emission HMM.
Args:
x: (T,) integer observations in {0, ..., V-1}
K: number of hidden states
V: vocabulary size (number of emission symbols)
num_iters: EM iterations
Returns:
pi0 (K,), P (K, K), theta (K, V), ll_history
'''
torch.manual_seed(seed)
T = len(x)
# Random initialisation
pi0 = torch.ones(K) / K
P = torch.ones(K, K) / K
theta = torch.rand(K, V)
theta = theta / theta.sum(1, keepdim=True)
ll_history = []
for _ in range(num_iters):
# E-step
log_ll = cat_log_likes(x, theta.log())
log_alphas, log_norms = forward_pass(pi0.log(), P.log(), log_ll)
log_betas = backward_pass(P.log(), log_ll)
gamma = posterior_marginals(log_alphas, log_betas, log_ll) # (T, K)
xi = posterior_pairs(log_alphas, log_betas, log_ll, P.log()) # (T-1,K,K)
ll_history.append(log_norms.sum().item())
# M-step
pi0 = (gamma[0] + 1e-8) / (gamma[0] + 1e-8).sum()
xi_sum = xi.sum(0) + 1e-8 # (K, K)
P = xi_sum / xi_sum.sum(1, keepdim=True)
# Weighted counts for each emission symbol
x_oh = torch.zeros(T, V).scatter_(1, x.unsqueeze(1), 1.0) # (T, V) one-hot
counts = gamma.T @ x_oh + 1e-8 # (K, V)
theta = counts / counts.sum(1, keepdim=True)
return pi0, P, theta, ll_history
Source
# ── Simulate from the dishonest casino ───────────────────────────────────────
K_true = 2 # states: 0=fair, 1=loaded
V = 6 # die faces 0..5
T_sim = 300
pi0_true = torch.tensor([1.0, 0.0])
P_true = torch.tensor([[0.95, 0.05],
[0.10, 0.90]])
theta_true = torch.tensor([[1/6]*6,
[0.1, 0.1, 0.1, 0.1, 0.1, 0.5]])
torch.manual_seed(42)
z_true = torch.zeros(T_sim, dtype=torch.long)
x_obs = torch.zeros(T_sim, dtype=torch.long)
z_true[0] = dist.Categorical(pi0_true).sample()
x_obs[0] = dist.Categorical(theta_true[z_true[0]]).sample()
for t in range(1, T_sim):
z_true[t] = dist.Categorical(P_true[z_true[t-1]]).sample()
x_obs[t] = dist.Categorical(theta_true[z_true[t]]).sample()
# ── Run EM to learn parameters ───────────────────────────────────────────────
pi0_em, P_em, theta_em, ll_hist = em_hmm_cat(x_obs, K=2, V=6, num_iters=80)
# Align states (EM may swap labels)
# State with higher p(6) should be "loaded"
loaded = theta_em[:, 5].argmax().item()
fair = 1 - loaded
# Posterior marginals with learned parameters
log_ll_em = cat_log_likes(x_obs, theta_em.log())
la, ln = forward_pass(pi0_em.log(), P_em.log(), log_ll_em)
lb = backward_pass(P_em.log(), log_ll_em)
gamma_em = posterior_marginals(la, lb, log_ll_em)
# ── Plot ──────────────────────────────────────────────────────────────────────
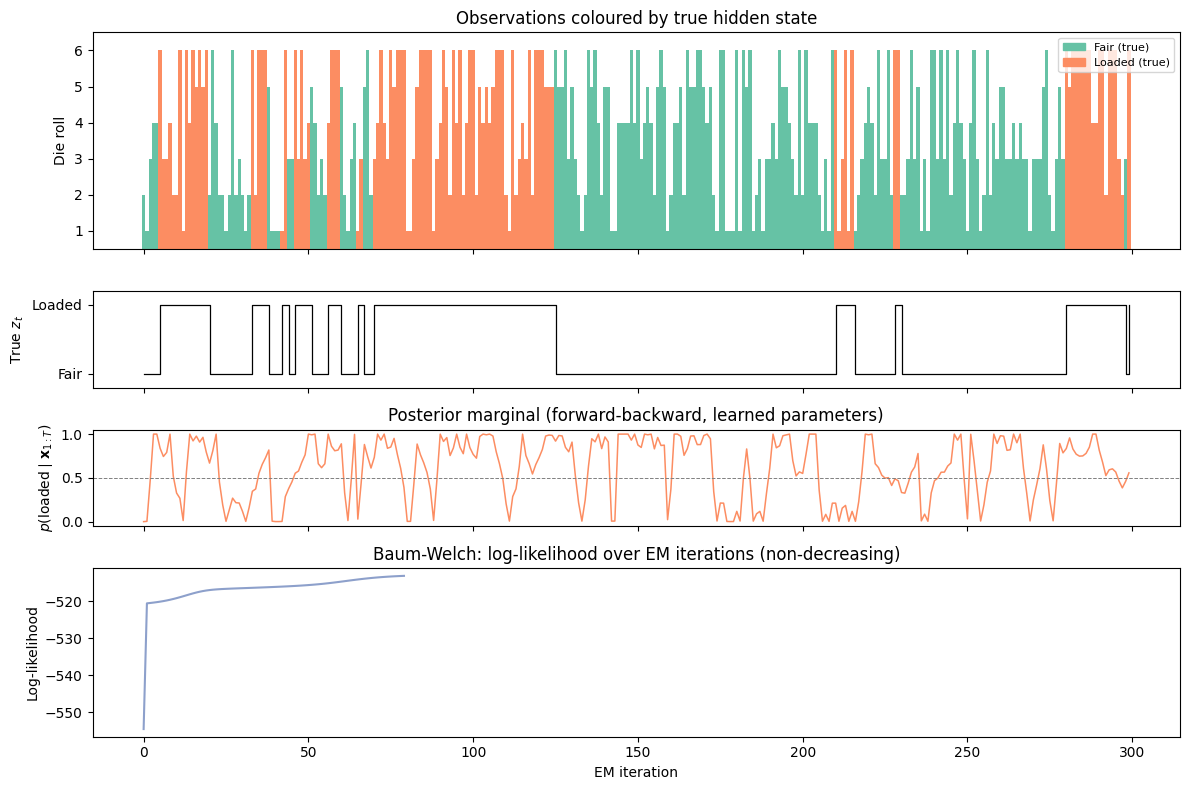
fig, axes = plt.subplots(4, 1, figsize=(12, 8), sharex=True,
gridspec_kw={'height_ratios': [1.8, 0.8, 0.8, 1.4]})
# Panel 1: die rolls coloured by true state
ax = axes[0]
colors = [palette[0] if z_true[t] == 0 else palette[1] for t in range(T_sim)]
ax.bar(range(T_sim), x_obs.numpy() + 1, color=colors, width=1.0, linewidth=0)
ax.set_ylabel('Die roll')
ax.set_ylim(0.5, 6.5)
ax.set_yticks([1, 2, 3, 4, 5, 6])
ax.set_title('Observations coloured by true hidden state')
fair_patch = mpatches.Patch(color=palette[0], label='Fair (true)')
loaded_patch = mpatches.Patch(color=palette[1], label='Loaded (true)')
ax.legend(handles=[fair_patch, loaded_patch], loc='upper right', fontsize=8)
# Panel 2: true hidden state sequence
ax = axes[1]
ax.step(range(T_sim), z_true.numpy(), where='post', color='k', lw=0.9)
ax.set_ylabel('True $z_t$')
ax.set_ylim(-0.2, 1.2)
ax.set_yticks([0, 1])
ax.set_yticklabels(['Fair', 'Loaded'])
# Panel 3: posterior p(loaded | all data) using learned parameters
ax = axes[2]
ax.plot(range(T_sim), gamma_em[:, loaded].numpy(), color=palette[1], lw=1.1)
ax.axhline(0.5, color='gray', linestyle='--', lw=0.7)
ax.set_ylabel('$p(\mathrm{loaded} \mid \mathbf{x}_{1:T})$')
ax.set_ylim(-0.05, 1.05)
ax.set_title('Posterior marginal (forward-backward, learned parameters)')
# Panel 4: ELBO / log-likelihood over EM iterations
ax = axes[3]
ax.plot(ll_hist, color=palette[2], lw=1.5)
ax.set_xlabel('EM iteration')
ax.set_ylabel('Log-likelihood')
ax.set_title('Baum-Welch: log-likelihood over EM iterations (non-decreasing)')
plt.tight_layout()
plt.show()
# Print learned vs true emission probabilities
print("Learned emission probs (fair die) :", theta_em[fair].numpy().round(3))
print("True emission probs (fair die) :", theta_true[0].numpy().round(3))
print()
print("Learned emission probs (loaded die):", theta_em[loaded].numpy().round(3))
print("True emission probs (loaded die):", theta_true[1].numpy().round(3))
print()
print(f"Learned transition matrix:\n{P_em.numpy().round(3)}")
print(f"True transition matrix:\n{P_true.numpy()}")
<>:63: SyntaxWarning: invalid escape sequence '\m'
<>:63: SyntaxWarning: invalid escape sequence '\m'
/tmp/ipykernel_2718/3219947313.py:63: SyntaxWarning: invalid escape sequence '\m'
ax.set_ylabel('$p(\mathrm{loaded} \mid \mathbf{x}_{1:T})$')
Learned emission probs (fair die) : [0.304 0.209 0.193 0. 0.02 0.274]
True emission probs (fair die) : [0.167 0.167 0.167 0.167 0.167 0.167]
Learned emission probs (loaded die): [0.002 0.101 0.149 0.236 0.201 0.311]
True emission probs (loaded die): [0.1 0.1 0.1 0.1 0.1 0.5]
Learned transition matrix:
[[0.703 0.297]
[0.188 0.812]]
True transition matrix:
[[0.95 0.05]
[0.1 0.9 ]]
Conclusion¶
| Component | Key idea |
|---|---|
| HMM | Mixture model with a Markov prior on |
| Forward pass | Recursive computation of |
| Backward pass | Recursive computation of |
| Forward-backward | Combines both to give |
| Baum-Welch / EM | E-step = forward-backward; M-step = weighted MLE |
Extensions worth knowing:
Viterbi algorithm: replaces with to find the most likely state sequence .
Autoregressive HMMs: each emission depends on both and recent observations Wiltschko et al., 2015.
Bayesian HMMs: place a Dirichlet prior on , rows of , and emission parameters; use Gibbs sampling or VI.
Infinite HMMs (iHMMs): nonparametric prior lets the number of states grow with the data.
- Wiltschko, A. B., Johnson, M. J., Iurilli, G., Peterson, R. E., Katon, J. M., Pashkovski, S. L., Abraira, V. E., Adams, R. P., & Datta, S. R. (2015). Mapping sub-second structure in mouse behavior. Neuron, 88(6), 1121–1135.
- Rabiner, L., & Juang, B. (1986). An introduction to hidden Markov models. Ieee Assp Magazine, 3(1), 4–16.