Directed graphical models (DGMs) provide a language for representing joint distributions over many variables compactly. By encoding conditional independence assumptions as a directed acyclic graph (DAG), we can specify and reason about high-dimensional distributions that would otherwise be intractable.
In this lecture we cover:
Product-rule factorization and the DAG representation
Plate notation for repeated structure
Conditional independence and the Markov blanket
Exchangeability and de Finetti’s theorem
Source
import io
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from daft import PGM
def pgm_to_img(pgm):
"""Render a PGM to an RGBA image array."""
pgm.render()
ax = plt.gca()
ax.set_aspect('equal')
buf = io.BytesIO()
fig = plt.gcf()
fig.savefig(buf, format='png', bbox_inches='tight', dpi=150)
buf.seek(0)
img = mpimg.imread(buf)
plt.close(fig)
return imgDirected Graphical Models¶
Motivation¶
Consider a joint distribution over discrete variables, each taking values in . An arbitrary distribution on such variables requires parameters — exponential in . Even for modest and this is enormous.
The product rule always lets us factor any joint distribution as a chain,
But this representation is equally expensive: conditioning on all preceding variables.
The key idea of graphical models is that many conditional independencies in real-world distributions make most of these conditions unnecessary. If only depends on a small subset of the preceding variables (its parents), then
The number of parameters is now , which is manageable when is small.
Definition¶
A directed graphical model (DGM), or Bayesian network, represents this factorization as a directed acyclic graph (DAG):
Each node corresponds to a variable — it may be discrete or continuous, scalar or multidimensional.
A directed edge from node to node means .
The graph must be acyclic (no directed cycles), which ensures the product factorization is well-defined.
The absence of an edge from to encodes a conditional independence assumption: does not depend directly on (given ’s other parents). The more edges we omit, the more compact the representation.
Source
# DAG for p(x1)p(x2)p(x3)p(x4|x1,x2,x3)p(x5|x1,x3)p(x6|x4)p(x7|x4,x5)
pgm = PGM()
# Row 0 (top): roots x1, x2, x3
pgm.add_node("x1", r"$x_1$", 0, 2)
pgm.add_node("x2", r"$x_2$", 2, 2)
pgm.add_node("x3", r"$x_3$", 4, 2)
# Row 1: x4, x5
pgm.add_node("x4", r"$x_4$", 1, 1)
pgm.add_node("x5", r"$x_5$", 3, 1)
# Row 2 (bottom): x6, x7
pgm.add_node("x6", r"$x_6$", 0, 0)
pgm.add_node("x7", r"$x_7$", 2, 0)
# Edges
for u, v in [("x1","x4"),("x2","x4"),("x3","x4"),
("x1","x5"),("x3","x5"),
("x4","x6"),("x4","x7"),("x5","x7")]:
pgm.add_edge(u, v)
pgm.render()
plt.gca().set_aspect('equal')
plt.show()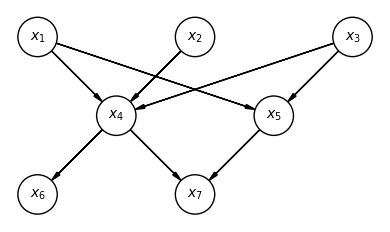
Plate Notation¶
Many probabilistic models have repeated structure — the same conditional distribution applied to many variables. For example, the diagonal-covariance Gaussian,
describes independent variables sharing the same conditional form. Drawing separate nodes would be redundant.
Plate notation compresses repeated structure by drawing a single representative node inside a rectangle (the plate), labelled with the index set. A node outside the plate is shared across all repetitions; a node inside is replicated once per index.
Plates can be nested: a plate labelled inside a plate labelled represents variables, one for each pair. We will use nested plates extensively in the hierarchical model below.
The figure below shows how a shared parameter generating i.i.d. observations is written in plate notation. Shaded nodes denote observed variables.
Source
# Plate notation: shared theta, N i.i.d. observed x_n
pgm = PGM()
pgm.add_node("theta", r"$\theta$", 1, 2)
pgm.add_node("xn", r"$x_n$", 1, 1, observed=True)
pgm.add_plate([0.4, 0.4, 1.2, 1.2], label=r"$n = 1, \ldots, N$",
position="bottom right")
pgm.add_edge("theta", "xn")
pgm.render()
plt.gca().set_aspect('equal')
plt.show()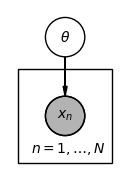
Conditional Independence¶
We write if and are conditionally independent given :
Conditional independencies in a DGM can be read off by examining three basic motifs (see Murphy or Bishop for the full d-separation rules):
Chain : (conditioning on the middle node blocks the path).
Fork : (conditioning on the common cause blocks the path).
Collider : (conditioning on a common effect opens a previously blocked path — often called explaining away).
Source
# Three d-separation motifs: chain, fork, collider
fig, axes = plt.subplots(1, 3, figsize=(9, 2.5))
for ax, (title, edges) in zip(axes, [
("Chain", [("xi","xk"),("xk","xj")]),
("Fork", [("xk","xi"),("xk","xj")]),
("Collider", [("xi","xk"),("xj","xk")]),
]):
pgm = PGM()
pgm.add_node("xi", r"$x_i$", 0, 0)
pgm.add_node("xk", r"$x_k$", 1, 0)
pgm.add_node("xj", r"$x_j$", 2, 0)
for u, v in edges:
pgm.add_edge(u, v)
img = pgm_to_img(pgm)
ax.imshow(img)
ax.axis("off")
ax.set_title(title, fontsize=11)
plt.tight_layout()
plt.show()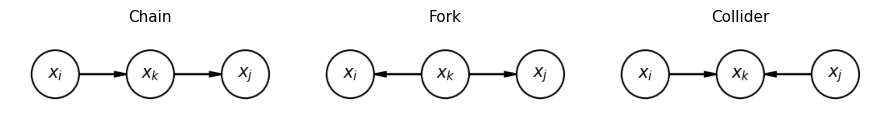
Markov Blanket¶
The Markov blanket of a node is the set of variables that, when conditioned on, render independent of all other variables. In a DGM, the Markov blanket of consists of:
’s parents,
’s children, and
the other parents of ’s children (the co-parents).
These are exactly the variables that appear alongside in at least one factor of the joint distribution. Given its Markov blanket, is conditionally independent of all remaining variables — a fact we will exploit repeatedly in deriving conditional posteriors.
Exchangeability and de Finetti’s Theorem¶
Suppose we want to model a collection of variables and have no information that distinguishes or orders them. A natural requirement is exchangeability: the joint distribution is invariant to any permutation ,
The simplest exchangeable distribution assumes the variables are i.i.d., . More generally, we may assume the variables are conditionally independent given a latent parameter which is marginalized over,
Marginally, the are not independent — observing some updates our belief about and therefore about other . But they are exchangeable.
De Finetti’s theorem states that as , any suitably well-behaved exchangeable distribution on can be represented in exactly this mixture form. The theorem does not hold in general for finite , but it provides a compelling motivation for Bayesian hierarchical models: placing a prior over and conditioning on the data is the correct procedure when observations are exchangeable.
Conclusion¶
Directed graphical models provide a principled way to represent the conditional independence structure of a joint distribution. The key results are:
Any joint distribution factors as a product of conditionals over a DAG; absent edges encode conditional independence assumptions that reduce the number of parameters.
Plate notation compresses repeated structure and makes hierarchical models easy to read.
The Markov blanket of a node determines its complete conditional — the ingredient needed for Gibbs sampling (Lecture 5).
Exchangeability motivates placing a shared prior over group-level parameters, and de Finetti’s theorem gives this practice a theoretical foundation.
- Murphy, K. P. (2023). Probabilistic Machine Learning: Advanced Topics. MIT Press. https://probml.github.io/pml-book/book2.html
- Bishop, C. M. (2006). Pattern recognition and machine learning. Springer.