The previous chapter introduced Hidden Markov Models, where the hidden state was discrete. Here we keep the same graphical model but allow to be continuous. When both the dynamics and the emissions are Gaussian and linear, the model is called a Linear Dynamical System (LDS), also known as a linear Gaussian state space model. The key payoff is that the forward-backward algorithm reduces to a pair of closed-form matrix recursions: the Kalman filter Kalman, 1960 and the Rauch-Tung-Striebel (RTS) smoother Rauch et al., 1965.
Source
import torch
import torch.distributions as dist
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.patches import Ellipse
import numpy as np
palette = list(plt.cm.Set2.colors)
torch.manual_seed(305)
<torch._C.Generator at 0x7f7ff41bf9f0>From HMMs to State Space Models¶
An HMM uses the same graphical model as a general state space model:
In an HMM, is discrete and the forward-backward algorithm sweeps over values at each time step. Nothing in the graphical model requires discrete states, however. If is continuous, the sums in the forward-backward recursions become integrals. In general those integrals are intractable, but for the Gaussian linear case they have a closed form.
The Linear Dynamical System¶
The LDS is defined by Gaussian dynamics and Gaussian emissions:
| Symbol | Dimension | Role |
|---|---|---|
| Dynamics (transition) matrix | ||
| Dynamics bias | ||
| Dynamics noise covariance | ||
| Emission (observation) matrix | ||
| Emission bias | ||
| Emission noise covariance |
Special cases. Setting and recovers Probabilistic PCA (Chapter 6): the latent states are i.i.d. Gaussians. The LDS adds temporal dependence through the dynamics matrix .
Stability. The dynamics are stable iff all eigenvalues of satisfy . If any eigenvalue exceeds 1 in magnitude, the latent state diverges.
The Kalman Filter¶
The forward pass for the LDS is the Kalman filter, which computes the filtering distributions
via two alternating steps. Assuming the predicted distribution is in hand, we proceed as follows.
Update Step¶
Condition on the new observation . The joint distribution of given is jointly Gaussian:
Applying the conditional Gaussian formula (Chapter 2), the posterior is:
where
The Kalman gain interpolates between the prior prediction and the observation: when (perfect sensors) and we trust the data entirely; when (useless sensors) and we ignore the data.
Predict Step¶
Marginalise over to obtain the predicted distribution for time :
Base case: and .
Marginal Likelihood¶
As a byproduct, each update step gives the one-step predictive likelihood:
so .
The RTS Smoother¶
The Kalman filter only uses past observations; it produces the filtering distribution . The smoothing distribution uses all observations and is generally tighter.
Backward Recursion¶
By the Markov property, is conditionally independent of given . This gives:
Marginalising over :
For the Gaussian LDS all three factors are Gaussian and the integral is tractable. The result is the Rauch-Tung-Striebel (RTS) smoother:
with backward recursion (starting from , from the filter):
The smoother gain plays the same role as the Kalman gain: it determines how much the smoothed estimate at time is adjusted based on new information from time .
Information Form and Sparse Linear Algebra¶
There is an elegant alternative view. Write the joint posterior as:
Expanding and collecting, the exponent is a quadratic form in , so the posterior is a multivariate Gaussian with natural parameters (precision matrix) and .
The key structural observation is that is block-tridiagonal: only the diagonal blocks and the off-diagonal blocks are non-zero, reflecting the Markov structure. The posterior mean can therefore be computed in rather than the naive by exploiting the block-tridiagonal sparsity — and this block-tridiagonal solve is exactly equivalent to the Kalman filter and RTS smoother.
Beyond the Linear Gaussian Case¶
For nonlinear dynamics or non-Gaussian noise, the Kalman filter no longer applies exactly. Common approximations include:
| Method | Idea |
|---|---|
| Extended Kalman Filter (EKF) | Linearise around the current estimate |
| Unscented Kalman Filter (UKF) | Propagate a set of sigma points through |
| Sequential Monte Carlo (SMC) | Represent by weighted particles |
Sequential Monte Carlo. The forward messages are proportional to the predictive distributions . We can approximate this distribution with weighted particles :
Sample (propagate particles).
Weight (score against observation).
Normalise and resample with replacement according to .
The marginal likelihood is approximated by .
# ── Kalman filter ─────────────────────────────────────────────────────────────
def kalman_filter(y, A, b, Q, C, d, R, mu0, Sigma0):
'''Kalman filter for a linear Gaussian SSM.
Model:
z_1 ~ N(mu0, Sigma0)
z_t|z_{t-1} ~ N(A z_{t-1} + b, Q)
x_t|z_t ~ N(C z_t + d, R)
Args:
y: (T, Dx) observations
A: (Dz, Dz) dynamics matrix
b: (Dz,) dynamics bias
Q: (Dz, Dz) dynamics noise covariance
C: (Dx, Dz) emission matrix
d: (Dx,) emission bias
R: (Dx, Dx) emission noise covariance
mu0: (Dz,) initial state mean
Sigma0: (Dz, Dz) initial state covariance
Returns:
filtered_means: (T, Dz) mu_{t|t}
filtered_covs: (T, Dz, Dz) Sigma_{t|t}
predicted_means: (T, Dz) mu_{t|t-1}
predicted_covs: (T, Dz, Dz) Sigma_{t|t-1}
log_marginal_likelihood: scalar
'''
T, Dx = y.shape
Dz = mu0.shape[0]
I = torch.eye(Dz)
filtered_means = torch.zeros(T, Dz)
filtered_covs = torch.zeros(T, Dz, Dz)
predicted_means = torch.zeros(T, Dz)
predicted_covs = torch.zeros(T, Dz, Dz)
log_ml = 0.0
mu_pred = mu0.clone()
Sigma_pred = Sigma0.clone()
for t in range(T):
# Store predicted distribution p(z_t | x_{1:t-1})
predicted_means[t] = mu_pred
predicted_covs[t] = Sigma_pred
# ── Update: condition on x_t ──────────────────────────────────
innov = y[t] - C @ mu_pred - d # innovation (Dx,)
S = C @ Sigma_pred @ C.T + R # innov. cov (Dx, Dx)
K = Sigma_pred @ C.T @ torch.linalg.inv(S) # Kalman gain (Dz, Dx)
mu_filt = mu_pred + K @ innov
Sigma_filt = (I - K @ C) @ Sigma_pred
# Marginal likelihood contribution log N(x_t | C mu_{t|t-1} + d, S)
log_ml += dist.MultivariateNormal(C @ mu_pred + d, S).log_prob(y[t])
filtered_means[t] = mu_filt
filtered_covs[t] = Sigma_filt
# ── Predict: propagate to t+1 ─────────────────────────────────
if t < T - 1:
mu_pred = A @ mu_filt + b
Sigma_pred = A @ Sigma_filt @ A.T + Q
return filtered_means, filtered_covs, predicted_means, predicted_covs, log_ml
# ── RTS Smoother ──────────────────────────────────────────────────────────────
def rts_smoother(A, filtered_means, filtered_covs, predicted_means, predicted_covs):
'''Rauch-Tung-Striebel backward smoother.
Args:
A: (Dz, Dz) dynamics matrix
filtered_means: (T, Dz) from kalman_filter
filtered_covs: (T, Dz, Dz)
predicted_means: (T, Dz)
predicted_covs: (T, Dz, Dz)
Returns:
smoothed_means: (T, Dz) mu_{t|T}
smoothed_covs: (T, Dz, Dz) Sigma_{t|T}
'''
T = filtered_means.shape[0]
smoothed_means = filtered_means.clone()
smoothed_covs = filtered_covs.clone()
for t in range(T - 2, -1, -1):
# Smoother gain G_t = Sigma_{t|t} A^T Sigma_{t+1|t}^{-1}
G = filtered_covs[t] @ A.T @ torch.linalg.inv(predicted_covs[t + 1])
smoothed_means[t] = (filtered_means[t]
+ G @ (smoothed_means[t + 1] - predicted_means[t + 1]))
smoothed_covs[t] = (filtered_covs[t]
+ G @ (smoothed_covs[t + 1] - predicted_covs[t + 1]) @ G.T)
return smoothed_means, smoothed_covs
Source
# ── 2D tracking demo: constant-velocity model ─────────────────────────────────
# State: z = [x, y, vx, vy] (position + velocity in 2D)
# Observation: x = [px, py] (noisy position)
import numpy as np
dt = 0.4
T_demo = 60
torch.manual_seed(42)
# Model parameters
A = torch.eye(4)
A[0, 2] = dt
A[1, 3] = dt
b = torch.zeros(4)
Q = torch.diag(torch.tensor([1e-4, 1e-4, 0.05, 0.05]))
C = torch.zeros(2, 4)
C[0, 0] = 1.0
C[1, 1] = 1.0
d = torch.zeros(2)
R = 0.4 * torch.eye(2)
mu0 = torch.tensor([0.0, 0.0, 0.8, 0.3])
Sigma0 = 0.1 * torch.eye(4)
# ── Simulate from the model ───────────────────────────────────────────────────
z_true = torch.zeros(T_demo, 4)
y_obs = torch.zeros(T_demo, 2)
z_true[0] = dist.MultivariateNormal(mu0, Sigma0).sample()
y_obs[0] = dist.MultivariateNormal(C @ z_true[0] + d, R).sample()
for t in range(1, T_demo):
z_true[t] = dist.MultivariateNormal(A @ z_true[t-1] + b, Q).sample()
y_obs[t] = dist.MultivariateNormal(C @ z_true[t] + d, R).sample()
# ── Run filter and smoother ───────────────────────────────────────────────────
filt_mu, filt_S, pred_mu, pred_S, log_ml = kalman_filter(
y_obs, A, b, Q, C, d, R, mu0, Sigma0)
smooth_mu, smooth_S = rts_smoother(A, filt_mu, filt_S, pred_mu, pred_S)
print(f"Log marginal likelihood: {log_ml:.2f}")
# ── Helper: draw a 2-sigma covariance ellipse (numpy-based) ──────────────────
def cov_ellipse(ax, mean_np, cov_np, n_std=2.0, **kwargs):
from matplotlib.patches import Ellipse
eigenvalues, eigenvectors = np.linalg.eigh(cov_np)
angle = np.degrees(np.arctan2(eigenvectors[1, -1], eigenvectors[0, -1]))
width, height = 2 * n_std * np.sqrt(eigenvalues)
ell = Ellipse(xy=mean_np, width=width, height=height, angle=angle, **kwargs)
ax.add_patch(ell)
# ── Plot ──────────────────────────────────────────────────────────────────────
fig, axes = plt.subplots(1, 2, figsize=(13, 5))
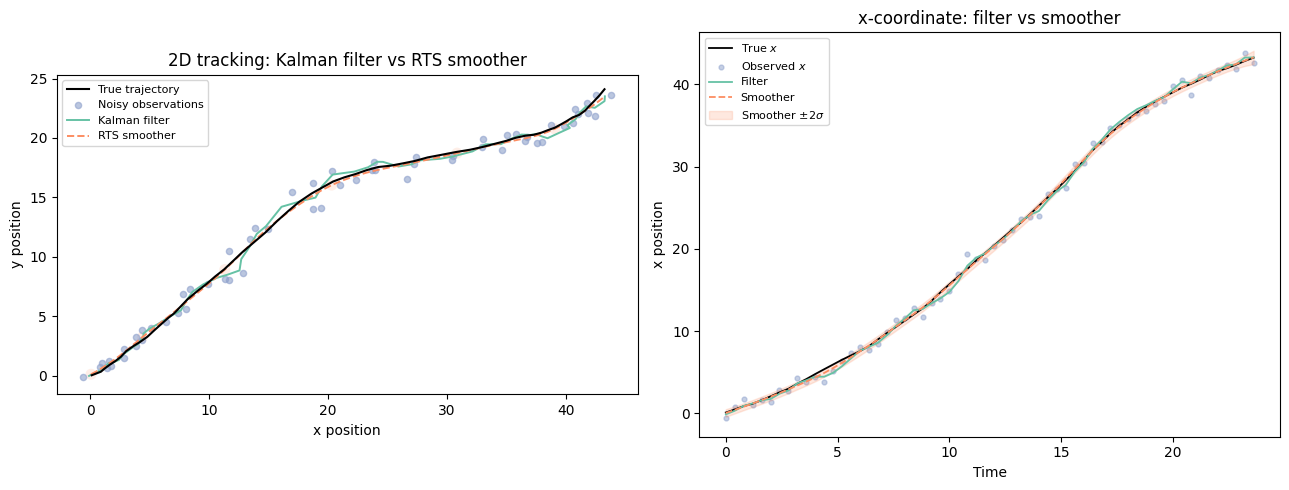
# ── Panel 1: 2D trajectories ──────────────────────────────────────────────────
ax = axes[0]
ax.plot(z_true[:, 0], z_true[:, 1], 'k-', lw=1.5, zorder=4, label='True trajectory')
ax.scatter(y_obs[:, 0], y_obs[:, 1], s=20, alpha=0.6,
color=palette[2], zorder=3, label='Noisy observations')
ax.plot(filt_mu[:, 0], filt_mu[:, 1],
color=palette[0], lw=1.4, zorder=3, label='Kalman filter')
ax.plot(smooth_mu[:, 0], smooth_mu[:, 1],
color=palette[1], lw=1.4, linestyle='--', zorder=3, label='RTS smoother')
for t in range(0, T_demo, 10):
cov_np = smooth_S[t, :2, :2].detach().numpy()
mean_np = smooth_mu[t, :2].detach().numpy()
cov_ellipse(ax, mean_np, cov_np,
n_std=2, facecolor=palette[1], alpha=0.15, edgecolor='none')
ax.set_title('2D tracking: Kalman filter vs RTS smoother')
ax.set_xlabel('x position')
ax.set_ylabel('y position')
ax.legend(fontsize=8)
ax.set_aspect('equal')
# ── Panel 2: x-coordinate time series ─────────────────────────────────────────
ax = axes[1]
t_vals = dt * np.arange(T_demo)
ax.plot(t_vals, z_true[:, 0].numpy(), 'k-', lw=1.3, label='True $x$')
ax.scatter(t_vals, y_obs[:, 0].numpy(), s=12, alpha=0.5,
color=palette[2], label='Observed $x$')
ax.plot(t_vals, filt_mu[:, 0].numpy(), color=palette[0], lw=1.3, label='Filter')
ax.plot(t_vals, smooth_mu[:, 0].numpy(), color=palette[1],
lw=1.3, linestyle='--', label='Smoother')
sig_x = smooth_S[:, 0, 0].sqrt().numpy()
ax.fill_between(t_vals,
smooth_mu[:, 0].numpy() - 2 * sig_x,
smooth_mu[:, 0].numpy() + 2 * sig_x,
alpha=0.2, color=palette[1], label='Smoother $\pm 2\sigma$')
ax.set_title('x-coordinate: filter vs smoother')
ax.set_xlabel('Time')
ax.set_ylabel('x position')
ax.legend(fontsize=8)
plt.tight_layout()
plt.show()
<>:93: SyntaxWarning: invalid escape sequence '\p'
<>:93: SyntaxWarning: invalid escape sequence '\p'
/tmp/ipykernel_2744/2116936060.py:93: SyntaxWarning: invalid escape sequence '\p'
alpha=0.2, color=palette[1], label='Smoother $\pm 2\sigma$')
Log marginal likelihood: -148.77
Conclusion¶
| Algorithm | Pass | Output | Cost |
|---|---|---|---|
| Kalman filter | Forward | ||
| RTS smoother | Backward |
The Kalman filter and RTS smoother are the exact continuous-state analogues of the discrete forward-backward algorithm: they exploit the Markov structure to compute marginals in linear time.
Learning. Given the smoothed marginals, the EM M-step for LDS parameters has closed forms analogous to the mixture model case. The sufficient statistics are , , and , all of which follow from the smoothed means and covariances. This is sometimes called the EM algorithm for LDS or the Shumway-Stoffer algorithm Shumway & Stoffer, 2000.
- Kalman, R. E. (1960). A new approach to linear filtering and prediction problems. Journal of Basic Engineering, 82(1), 35–45.
- Rauch, H. E., Striebel, C. T., & Tung, F. (1965). Maximum likelihood estimates of linear dynamic systems. AIAA Journal, 3(8), 1445–1450.
- Shumway, R. H., & Stoffer, D. S. (2000). Time Series Analysis and Its Applications. Springer.