Finite Bayesian mixture models require specifying the number of components in advance. Dirichlet process mixture models (DPMMs) are a Bayesian nonparametric alternative that allows the number of clusters to grow with the data. In this chapter we derive DPMMs from finite mixture models by taking , then study the random measure perspective and its connection to Poisson processes.
Source
import torch
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import t as student_t
Finite Bayesian Mixture Models¶
A finite Bayesian mixture model with components is defined by the following generative process:
Sample mixing proportions from a Dirichlet prior with concentration :
Sample component parameters:
Sample assignments and observations:
The joint distribution factorizes as:
Collapsing Out Parameters¶
When the likelihood is an exponential family and the prior is its conjugate,
we can marginalize (or collapse) the component parameters and mixture proportions in closed form.
After integrating out and , the marginal likelihood over assignments and observations is:
where the updated hyperparameters are:
and is the Dirichlet normalizing constant.
This is a general pattern: in conjugate exponential families, marginal likelihoods are ratios of posterior to prior normalizing functions.
Collapsed Gibbs Sampling¶
Working with the collapsed distribution enables efficient collapsed Gibbs sampling. The conditional distribution of , holding all other assignments fixed, simplifies to:
where is the number of other points in cluster .
The two factors have intuitive interpretations:
The cluster size term comes from the Dirichlet–Multinomial conjugacy: larger clusters attract new points.
The posterior predictive term measures how well fits with the other points in cluster .
The Dirichlet Process Mixture Model¶
Now set and take . In this limit, the collapsed Gibbs update simplifies to:
This is the DPMM collapsed Gibbs update. It is nonparametric — is never fixed, and the number of occupied clusters can grow or shrink at each step:
A cluster is created when a data point is assigned to a new cluster (probability controlled by ).
A cluster is deleted when its last data point is reassigned elsewhere.
The parameter is the concentration: larger creates more clusters.
The Chinese Restaurant Process (CRP)¶
The prior on partitions induced by the DPMM has a beautiful sequential description called the Chinese restaurant process (CRP). Imagine customers entering a restaurant one at a time:
Customer 1 sits at a new table.
Customer sits at an existing table with probability , or starts a new table with probability .
The resulting partition is the CRP prior on partitions of .
Stick-Breaking Construction¶
The DPMM can also be defined directly via a random measure:
where the weights follow a stick-breaking construction: sample and set
We write and call the concentration and the base measure.
def normal_log_posterior_predictive(x, cluster_points, mu0, kappa0, alpha0, beta0):
'''Log posterior predictive p(x | cluster_points) under Normal-NormalInverseGamma prior.
Model: x_n | mu, sigma^2 ~ N(mu, sigma^2)
mu | sigma^2 ~ N(mu0, sigma^2 / kappa0)
sigma^2 ~ IG(alpha0, beta0)
The marginal p(x | data) is a Student-t distribution.
Args:
x: new point, shape (D,)
cluster_points: tensor of shape (N_k, D); empty if prior predictive
mu0: prior mean, shape (D,)
kappa0: prior precision scaling (scalar)
alpha0: prior shape (scalar)
beta0: prior rate (scalar)
Returns:
log predictive density (scalar)
'''
D = x.shape[0]
n = len(cluster_points)
if n > 0:
x_bar = cluster_points.mean(0)
kappa_n = kappa0 + n
mu_n = (kappa0 * mu0 + n * x_bar) / kappa_n
alpha_n = alpha0 + n / 2.0
S = ((cluster_points - x_bar) ** 2).sum(0) # shape (D,)
diff = x_bar - mu0
beta_n = beta0 + 0.5 * S + 0.5 * kappa0 * n / kappa_n * diff ** 2
else:
kappa_n, mu_n, alpha_n, beta_n = kappa0, mu0, alpha0, beta0 * torch.ones(D)
# Marginal is a Student-t: t(2*alpha_n, mu_n, beta_n*(kappa_n+1)/(alpha_n*kappa_n))
df = 2.0 * alpha_n
scale2 = beta_n * (kappa_n + 1) / (alpha_n * kappa_n)
log_p = 0.0
for d in range(D):
log_p += float(student_t.logpdf(x[d].item(), df=df, loc=mu_n[d].item(),
scale=scale2[d].item() ** 0.5))
return log_p
def collapsed_gibbs_dpmm(X, alpha, mu0, kappa0, alpha0, beta0, num_iters=100, seed=0):
'''Collapsed Gibbs sampler for a DPMM with Normal-NormalInverseGamma components.
Args:
X: data, shape (N, D)
alpha: DP concentration
mu0, kappa0, alpha0, beta0: NIG prior hyperparameters
num_iters: number of Gibbs sweeps
seed: random seed
Returns:
z: final cluster assignments, shape (N,)
K_history: number of clusters at each iteration
'''
torch.manual_seed(seed)
N, D = X.shape
mu0 = mu0 * torch.ones(D)
# Initialize: one cluster per data point
z = torch.arange(N)
K_history = []
for it in range(num_iters):
for n in range(N):
# Remove n from its current cluster
z_n = z[n].item()
z[n] = -1
mask_n = (z >= 0)
unique_ks = z[mask_n].unique().tolist()
# Compute log probabilities for each existing cluster and a new one
log_probs = []
labels = []
for k in unique_ks:
pts = X[(z == k)]
N_k = len(pts)
log_like = normal_log_posterior_predictive(X[n], pts, mu0, kappa0, alpha0, beta0)
log_probs.append(np.log(N_k) + log_like)
labels.append(k)
# New cluster
log_like_new = normal_log_posterior_predictive(X[n], X[:0], mu0, kappa0, alpha0, beta0)
log_probs.append(np.log(alpha) + log_like_new)
labels.append(max(unique_ks + [-1]) + 1 if unique_ks else 0)
# Sample
log_probs = np.array(log_probs)
log_probs -= log_probs.max()
probs = np.exp(log_probs)
probs /= probs.sum()
chosen = np.random.choice(len(labels), p=probs)
z[n] = labels[chosen]
# Re-label clusters 0, 1, 2, ...
unique_labels = z.unique().tolist()
remap = {old: new for new, old in enumerate(unique_labels)}
z = torch.tensor([remap[zi.item()] for zi in z])
K_history.append(len(unique_labels))
return z, K_history
Source
torch.manual_seed(1)
np.random.seed(1)
# Generate data from 3 Gaussians
true_means = torch.tensor([[-2.0, 0.0], [2.0, 0.0], [0.0, 2.5]])
true_std = 0.6
N_per = 40
X_list = [true_means[k] + true_std * torch.randn(N_per, 2) for k in range(3)]
X = torch.cat(X_list)
N = len(X)
# Prior hyperparameters
mu0 = torch.zeros(2)
kappa0 = 0.5
alpha0 = 2.0
beta0 = 1.0
alpha_dp = 2.0
z_final, K_hist = collapsed_gibbs_dpmm(X, alpha=alpha_dp, mu0=mu0, kappa0=kappa0,
alpha0=alpha0, beta0=beta0,
num_iters=80, seed=42)
fig, axes = plt.subplots(1, 2, figsize=(11, 4))
# Left: final clustering
K_final = z_final.max().item() + 1
colors = plt.cm.tab10(np.linspace(0, 1, max(K_final, 10)))
for k in range(K_final):
mask = (z_final == k).numpy()
axes[0].scatter(X[mask, 0], X[mask, 1], c=[colors[k]], s=20, alpha=0.8)
for km, mu in enumerate(true_means):
axes[0].scatter(*mu, marker='*', s=200, c='k', zorder=5)
axes[0].set_title(f'DPMM clustering ($\\alpha={alpha_dp}$): {K_final} clusters found')
axes[0].set_xlabel('$x_1$'); axes[0].set_ylabel('$x_2$')
# Right: K over iterations
axes[1].plot(K_hist, color='tab:blue')
axes[1].axhline(3, ls='--', color='k', label='True K=3')
axes[1].set_xlabel('Gibbs iteration')
axes[1].set_ylabel('Number of clusters')
axes[1].set_title('Cluster count over iterations')
axes[1].legend()
plt.tight_layout()
plt.savefig('dpmm_demo.png', dpi=100, bbox_inches='tight')
plt.show()
print(f"Final number of clusters: {K_final}")
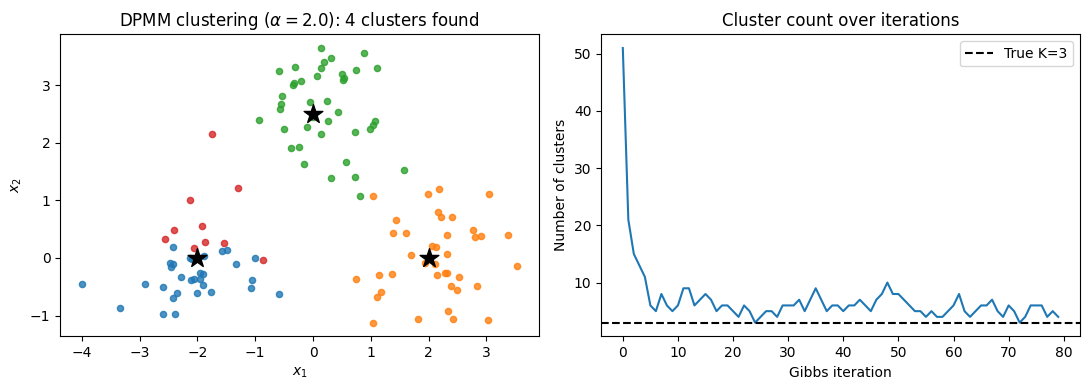
Final number of clusters: 4
Extensions and Related Models¶
Pitman–Yor Process¶
The Pitman–Yor process Orbanz, 2014 generalizes the DP by adding a discount parameter . We write where the weights use the modified stick-breaking:
When we recover the DP. When , the cluster sizes follow a power law, making the PYP well-suited to linguistic data where word frequencies exhibit Zipfian behavior.
The CRP analogue assigns customer to existing table with probability or to a new table with probability , where is the current number of tables.
Mixture of Finite Mixtures¶
DPMMs are often used to select automatically, but the DP almost surely generates infinitely many clusters as . When the true number of clusters is finite but unknown, mixture of finite mixture models (MFMMs) Miller & Harrison, 2018 are more appropriate:
A natural choice is . Collapsed Gibbs samplers for MFMMs have a similar form to the DPMM sampler but converge to a finite .
Poisson Random Measures¶
Dirichlet processes and Poisson processes are deeply connected. In fact, DPs are instances of completely random measures (CRMs) constructed from Poisson processes.
Random Measures¶
A random measure on is a measure-valued random variable. An atomic random measure takes the form:
where the weights and locations are random.
Poisson Random Measures¶
Construct a random measure by sampling weight–location pairs from a marked Poisson process on :
For a homogeneous measure the intensity factors: , where is a density on (the base measure ).
The Gamma Process and Dirichlet Process¶
Choose the weight intensity
Then , so has infinitely many atoms. Nevertheless, the total mass is almost surely finite. This is the gamma process .
Normalizing gives the Dirichlet process:
This is a key result: Dirichlet processes are normalized gamma processes.
Other Completely Random Measures¶
Different weight intensities yield other useful random measures:
| Weight intensity | Name |
|---|---|
| Gamma process → DP (after normalization) | |
| Stable process | |
| Beta process |
A CRM has the remarkable property that is independent of if and only if is a gamma process — the DP is the unique normalized CRM with this independence property Orbanz, 2014.
- Orbanz, P. (2014). Lecture notes on Bayesian nonparametrics. http://www.gatsby.ucl.ac.uk/~porbanz/papers/porbanz_BNP_draft.pdf
- Miller, J. W., & Harrison, M. T. (2018). Mixture models with a prior on the number of components. Journal of the American Statistical Association, 113(521), 340–356.