Gibbs sampling and random-walk Metropolis–Hastings are powerful, but they scale poorly with dimension. A random-walk proposal explores parameter space by diffusing, which requires steps to produce an independent sample in a -dimensional space.
Hamiltonian Monte Carlo (HMC) uses gradient information to make large, directed moves through parameter space, reducing that cost to roughly . The key idea is to interpret the negative log-probability as a potential energy surface and simulate the physics of a frictionless particle sliding over it.
In this chapter we:
Motivate gradient-based proposals starting from the Metropolis-Adjusted Langevin Algorithm (MALA)
Derive Hamilton’s equations and their key properties
Implement the leapfrog integrator, the numerical engine of HMC
Assemble the full HMC algorithm and verify it on a 2D correlated Gaussian
Compare HMC against random-walk MH as dimension grows
Source
import torch
from torch.distributions import MultivariateNormal
from pyro.ops.stats import effective_sample_size
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
/opt/hostedtoolcache/Python/3.13.13/x64/lib/python3.13/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Why Random Walks Are Slow¶
In Metropolis–Hastings with a symmetric Gaussian proposal , the chain diffuses through . To maintain a reasonable acceptance rate, the proposal s.d. must be comparable to the smallest marginal s.d. of the posterior. Each step is then roughly uncorrelated only in that most-constrained direction, while in wider directions many steps are needed to traverse the distribution. The upshot: gradient evaluations to produce one effectively independent sample Neal, 2011.
Metropolis-Adjusted Langevin Algorithm¶
One improvement is to drift the proposal toward high-probability regions using the gradient:
This Metropolis-Adjusted Langevin Algorithm (MALA) is a discrete-time approximation to the Langevin diffusion SDE. It needs only gradient evaluations per independent sample — better than random-walk MH, but we can do better still by taking multiple gradient steps per proposal. That is the idea behind HMC.
Hamiltonian Dynamics¶
Setup¶
Think of the negative log-probability as a potential energy surface. Introduce an auxiliary momentum variable (one per parameter) and define the Hamiltonian:
where is a symmetric positive-definite mass matrix (typically ).
The joint distribution over position and momentum factorizes:
Marginalizing out recovers the posterior, so samples of from this joint distribution are posterior samples.
Hamilton’s Equations¶
The state evolves in continuous time according to:
Three key properties of these dynamics make them useful for MCMC:
Reversibility: negating reverses the trajectory, making the proposal symmetric after a final momentum flip.
Conservation of energy: , so the Hamiltonian is constant along trajectories. If the dynamics could be simulated exactly, every HMC proposal would be accepted.
Volume preservation: the divergence of the vector field is zero (since ), so Liouville’s theorem applies and there is no Jacobian correction in the acceptance probability.
1D Example: Harmonic Oscillator¶
When , , and , Hamilton’s equations become , — a rotation in space. The exact solution is where , tracing a circle of constant Hamiltonian.
The Leapfrog Integrator¶
Euler vs. Leapfrog¶
Naïve Euler integration updates position and momentum simultaneously:
Euler integration does not preserve volume and introduces systematic energy drift even for small .
The leapfrog integrator interleaves half-steps for momentum with full steps for position:
Each update is a shear transformation (only some variables change, by amounts depending on the others), which has unit Jacobian. Composing shears preserves volume exactly. consecutive leapfrog steps cost gradient evaluations ( full steps sandwiched between two half-steps).
def leapfrog(q, p, grad_U, ε, L, M_inv):
"""Simulate L leapfrog steps of size ε.
Args:
q: (D,) position
p: (D,) momentum
grad_U: callable, returns gradient of U at q
ε: step size
L: number of leapfrog steps
M_inv: (D, D) inverse mass matrix
Returns:
q, p after L steps
"""
p = p - (ε / 2) * grad_U(q)
for _ in range(L - 1):
q = q + ε * (M_inv @ p)
p = p - ε * grad_U(q)
q = q + ε * (M_inv @ p)
p = p - (ε / 2) * grad_U(q)
return q, pSource
# Compare Euler and Leapfrog on a 1D harmonic oscillator: U(q)=q²/2, m=1
# Exact solution: (q,p) traces a circle of radius sqrt(q0²+p0²)
grad_U_1d = lambda q: q # dU/dq = q
M_inv_1d = torch.eye(1)
q0, p0 = torch.tensor([1.0]), torch.tensor([0.0])
steps = 30
# Euler
q_e, p_e = q0.clone(), p0.clone()
traj_euler = [(q_e.item(), p_e.item())]
for _ in range(steps):
p_e = p_e - 0.3 * q_e
q_e = q_e + 0.3 * p_e
traj_euler.append((q_e.item(), p_e.item()))
# Leapfrog (each call = 1 leapfrog step)
q_l, p_l = q0.clone(), p0.clone()
traj_leap = [(q_l.item(), p_l.item())]
for _ in range(steps):
q_l, p_l = leapfrog(q_l, p_l, grad_U_1d, 0.3, 1, M_inv_1d)
traj_leap.append((q_l.item(), p_l.item()))
# Exact circle
θ_circ = torch.linspace(0, 2 * torch.pi, 200)
q_circ, p_circ = torch.cos(θ_circ).numpy(), torch.sin(θ_circ).numpy()
fig, axs = plt.subplots(1, 2, figsize=(9, 4))
for ax, traj, title in [
(axs[0], traj_euler, 'Euler (ε = 0.3, 30 steps)'),
(axs[1], traj_leap, 'Leapfrog (ε = 0.3, 30 steps)')]:
qs, ps = zip(*traj)
ax.plot(q_circ, p_circ, 'k--', lw=1, label='exact')
ax.plot(qs, ps, 'o-', ms=4, label='numerical')
ax.set_xlabel('q'); ax.set_ylabel('p')
ax.set_title(title); ax.legend(); ax.set_aspect('equal')
plt.tight_layout()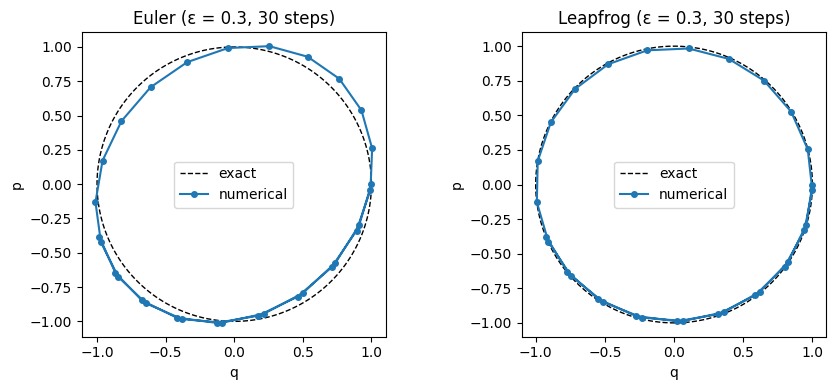
The HMC Algorithm¶
Using Hamiltonian Dynamics as a Proposal¶
HMC is Metropolis–Hastings on the joint distribution with proposals generated by leapfrog integration.
Algorithm (one HMC iteration starting from ):
Resample momentum: .
Leapfrog steps: integrate Hamilton’s equations for time to obtain proposed .
Flip momentum: (makes the proposal symmetric).
Accept/reject:
If were exactly conserved (exact dynamics), always. In practice the leapfrog introduces energy error, so some rejections occur, and a smaller gives a higher acceptance rate at the cost of smaller steps.
Tuning¶
The two hyperparameters are the step size and the number of steps :
controls accuracy: smaller → higher acceptance rate but smaller moves.
controls how far each proposal travels: larger → more decorrelated samples but more gradient evaluations per step.
A target acceptance rate of ~0.65 is commonly used for step-size adaptation. The No-U-Turn Sampler (NUTS; Hoffman & Gelman, 2014) automatically adapts by continuing the trajectory until it starts turning back on itself — eliminating the need to tune by hand. NUTS is the default sampler in Stan and Pyro.
def hmc(q0, log_joint, grad_U, ε, L, M, num_samples):
"""Hamiltonian Monte Carlo sampler.
Args:
q0: (D,) initial position
log_joint: callable: q -> scalar log p(q, X)
grad_U: callable: q -> (D,) gradient of -log_joint
ε: leapfrog step size
L: number of leapfrog steps per proposal
M: (D, D) mass matrix
num_samples: number of HMC iterations
Returns:
(num_samples+1, D) tensor of samples (including q0),
acceptance rate
"""
M_inv = torch.linalg.inv(M)
M_chol = torch.linalg.cholesky(M)
q = q0.clone()
samples = [q.clone()]
n_accept = 0
for _ in range(num_samples):
p = M_chol @ torch.randn_like(q)
H_curr = -log_joint(q) + 0.5 * p @ M_inv @ p
q_prop, p_prop = leapfrog(q.clone(), p.clone(), grad_U, ε, L, M_inv)
H_prop = -log_joint(q_prop) + 0.5 * p_prop @ M_inv @ p_prop
if torch.rand(1).item() < torch.exp(H_curr - H_prop).item():
q = q_prop
n_accept += 1
samples.append(q.clone())
return torch.stack(samples), n_accept / num_samplesDemo: 2D Correlated Gaussian¶
We test HMC on a correlated bivariate Gaussian with , whose contours form elongated ellipses. This is a hard case for random-walk MH: the optimal proposal must be tiny in the narrow direction, causing slow exploration of the wide direction.
torch.manual_seed(305)
# 2D correlated Gaussian: Σ with off-diagonal ρ
ρ = 0.95
Σ = torch.tensor([[1.0, ρ], [ρ, 1.0]])
Σ_inv = torch.linalg.inv(Σ)
def log_joint_2d(q):
return -0.5 * q @ Σ_inv @ q
def grad_U_2d(q):
return Σ_inv @ q # gradient of U = -log_joint
# Run HMC
M_2d = torch.eye(2)
q0_2d = torch.zeros(2)
samples_hmc, acc_hmc = hmc(q0_2d, log_joint_2d, grad_U_2d,
ε=0.2, L=20, M=M_2d, num_samples=2000)
print(f'HMC acceptance rate: {acc_hmc:.2f}')HMC acceptance rate: 0.99
Source
# Draw one HMC trajectory to illustrate the leapfrog path
torch.manual_seed(1)
q_demo = torch.tensor([-1.5, 1.0])
p_demo = M_2d @ torch.randn(2)
traj_q = [q_demo.clone()]
q_t, p_t = q_demo.clone(), p_demo.clone()
M_inv_2d = torch.linalg.inv(M_2d)
for _ in range(20):
q_t, p_t = leapfrog(q_t, p_t, grad_U_2d, 0.2, 1, M_inv_2d)
traj_q.append(q_t.clone())
traj_q = torch.stack(traj_q)
# Contour grid
g = torch.linspace(-3, 3, 200)
Q1, Q2 = torch.meshgrid(g, g, indexing='ij')
Z_grid = torch.stack([Q1.flatten(), Q2.flatten()], dim=1)
log_p = (-0.5 * (Z_grid @ Σ_inv * Z_grid).sum(1)).reshape(200, 200)
fig, axs = plt.subplots(1, 2, figsize=(11, 4.5))
# Left: trajectory
axs[0].contour(Q1.numpy(), Q2.numpy(), log_p.exp().numpy(), levels=6, colors='grey', linewidths=0.8)
axs[0].plot(traj_q[:, 0].numpy(), traj_q[:, 1].numpy(), 'b-o', ms=4, label='leapfrog path')
axs[0].plot(*traj_q[0].numpy(), 'go', ms=8, label='start')
axs[0].plot(*traj_q[-1].numpy(), 'rs', ms=8, label='proposal')
axs[0].set_title('One HMC leapfrog trajectory'); axs[0].legend()
axs[0].set_xlabel(r'$q_1$'); axs[0].set_ylabel(r'$q_2$')
# Right: scatter of samples
burnin = 200
axs[1].contour(Q1.numpy(), Q2.numpy(), log_p.exp().numpy(), levels=6, colors='grey', linewidths=0.8)
axs[1].scatter(samples_hmc[burnin:, 0].numpy(), samples_hmc[burnin:, 1].numpy(),
s=4, alpha=0.4, label='HMC samples')
axs[1].set_title('HMC samples (ε=0.2, L=20)')
axs[1].set_xlabel(r'$q_1$'); axs[1].set_ylabel(r'$q_2$')
plt.tight_layout()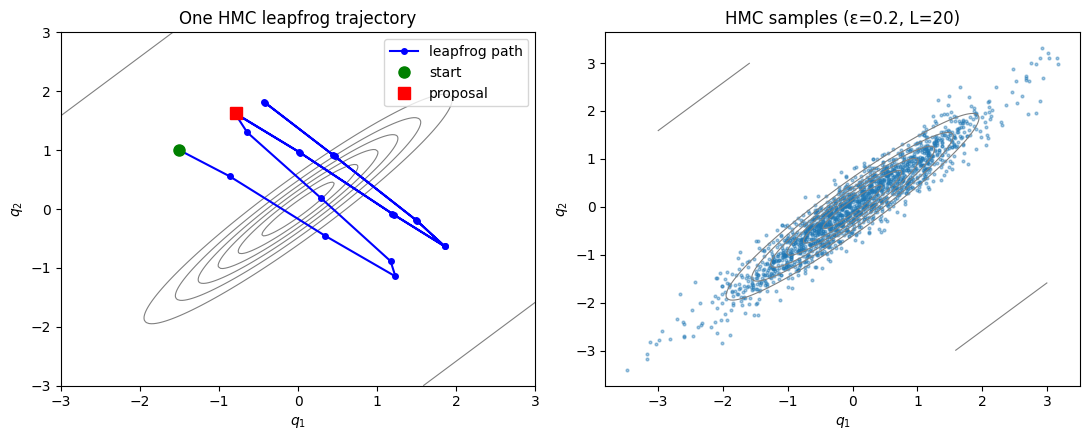
Comparison with Random-Walk MH¶
For random-walk MH on the same target, the proposal s.d. must be small enough to not overshoot the narrow direction (), which makes exploration of the wide direction slow.
# Random-walk MH on same 2D Gaussian
def rwmh(q0, log_joint, σ_prop, num_samples):
D = q0.shape[0]
q = q0.clone()
samples = [q.clone()]
n_accept = 0
for _ in range(num_samples):
q_prop = q + σ_prop * torch.randn(D)
log_α = log_joint(q_prop) - log_joint(q)
if torch.rand(1).item() < torch.exp(log_α).item():
q = q_prop
n_accept += 1
samples.append(q.clone())
return torch.stack(samples), n_accept / num_samples
samples_rw, acc_rw = rwmh(q0_2d, log_joint_2d, σ_prop=0.3, num_samples=2000)
print(f'RW-MH acceptance rate: {acc_rw:.2f}')
ess_hmc = effective_sample_size(samples_hmc[None, burnin:, 0]).item()
ess_rw = effective_sample_size(samples_rw[None, burnin:, 0]).item()
print(f'ESS q₁ — HMC: {ess_hmc:.0f}, RW-MH: {ess_rw:.0f} (out of {2000-burnin} samples)')RW-MH acceptance rate: 0.60
ESS q₁ — HMC: 2939, RW-MH: 18 (out of 1800 samples)
Source
fig, axs = plt.subplots(2, 2, figsize=(10, 6), sharex='col')
for col, (samps, label) in enumerate([(samples_hmc, 'HMC'), (samples_rw, 'RW-MH')]):
axs[0, col].plot(samps[burnin:burnin+500, 0].numpy())
axs[0, col].set_title(f'{label}: trace of $q_1$')
axs[0, col].set_ylabel(r'$q_1$')
axs[1, col].plot(samps[burnin:burnin+500, 1].numpy())
axs[1, col].set_ylabel(r'$q_2$')
axs[1, col].set_xlabel('Iteration')
plt.tight_layout()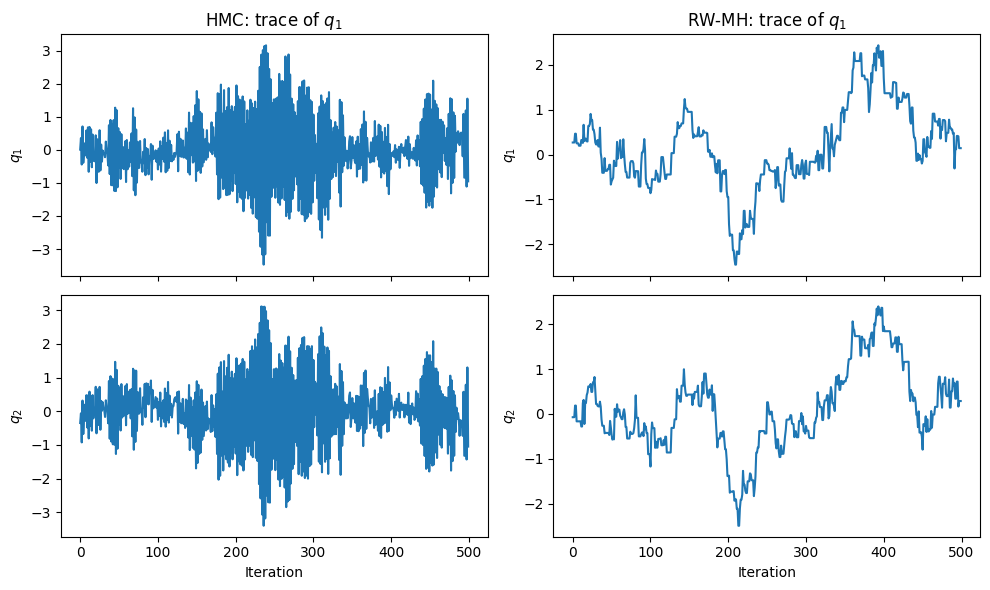
Scaling with Dimension¶
Benefits of Avoiding Random Walks¶
For a -dimensional isotropic Gaussian target the cost of obtaining one independent sample scales as:
| Algorithm | Gradient evaluations |
|---|---|
| Random-walk MH | |
| MALA | |
| HMC |
The advantage of HMC grows with the condition number of the posterior covariance. Random-walk MH must step at the scale of the most constrained dimension, requiring steps in the widest direction (where is the condition number). HMC integrates the gradient to traverse the wide direction efficiently in leapfrog steps.
Demonstration in Higher Dimensions¶
We compare HMC and random-walk MH on a -dimensional diagonal Gaussian with eigenvalues linearly spaced from 0.1 to 1 (condition number ).
torch.manual_seed(305)
results = {}
for D in [2, 10, 50, 100]:
λ_vec = torch.linspace(0.1, 1.0, D)
Σ_inv_d = torch.diag(1.0 / λ_vec)
def log_joint_d(q, Σ_inv=Σ_inv_d):
return -0.5 * q @ Σ_inv @ q
def grad_U_d(q, Σ_inv=Σ_inv_d):
return Σ_inv @ q
# HMC: step size ≈ smallest std, L = 20
ε_hmc = 0.8 * λ_vec.min().sqrt().item()
samp_h, _ = hmc(torch.zeros(D), log_joint_d, grad_U_d,
ε=ε_hmc, L=20, M=torch.eye(D), num_samples=2000)
ess_h = effective_sample_size(samp_h[None, 500:, 0]).item()
# RW-MH: proposal std ≈ smallest std
samp_r, _ = rwmh(torch.zeros(D), log_joint_d, σ_prop=λ_vec.min().sqrt().item(), num_samples=2000)
ess_r = effective_sample_size(samp_r[None, 500:, 0]).item()
results[D] = (ess_h, ess_r)
print(f'D={D:3d}: ESS HMC={ess_h:6.0f}, ESS RW-MH={ess_r:6.0f}')D= 2: ESS HMC= 8182, ESS RW-MH= 172
D= 10: ESS HMC= 6103, ESS RW-MH= 92
D= 50: ESS HMC= 5127, ESS RW-MH= 26
D=100: ESS HMC= 3838, ESS RW-MH= nan
Source
dims = sorted(results)
ess_h_vals = [results[d][0] for d in dims]
ess_r_vals = [results[d][1] for d in dims]
plt.figure(figsize=(6, 4))
plt.plot(dims, ess_h_vals, 's-', label='HMC')
plt.plot(dims, ess_r_vals, 'o-', label='RW-MH')
plt.xlabel('Dimension $D$')
plt.ylabel('ESS (out of 1500 post-burnin samples)')
plt.title('HMC vs. RW-MH effective sample size')
plt.legend()
plt.tight_layout()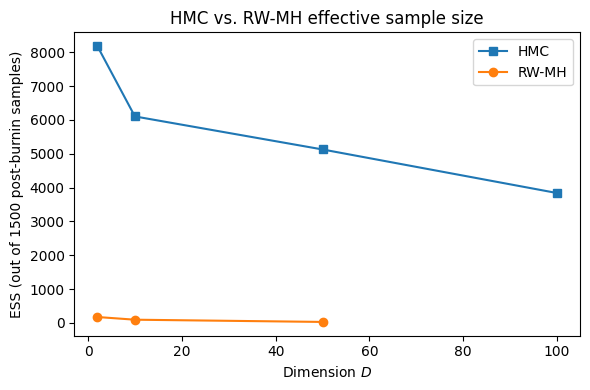
Practical Considerations¶
Step-Size Adaptation¶
The step size must be tuned:
Too large → large energy errors → low acceptance rate.
Too small → small moves → slow mixing.
A target acceptance rate of ~0.65 is recommended by Neal, 2011. A simple strategy during warm-up: increase when the acceptance rate exceeds the target, decrease when below. The dual averaging scheme of Hoffman & Gelman, 2014 provides a principled adaptive version used by default in Pyro and Stan.
No-U-Turn Sampler¶
Choosing by hand is also inconvenient. The No-U-Turn Sampler (NUTS) Hoffman & Gelman, 2014 extends the trajectory dynamically until it begins turning back (the inner product ), then selects a sample from along the trajectory using slice sampling. NUTS:
Eliminates the need to tune .
Adapts automatically during warm-up.
Is the default backend for Stan, PyMC, and Pyro’s
NUTSkernel.
An interactive visualization of HMC and NUTS:
https://
Conclusion¶
HMC exploits gradient information to make large, coherent moves through parameter space, vastly outperforming random-walk methods in high dimensions. Key points:
The potential energy is ; auxiliary momentum variables complete the Hamiltonian system.
Hamiltonian dynamics are reversible, energy-conserving, and volume-preserving — exactly the properties needed for a valid MCMC kernel.
The leapfrog integrator preserves these properties approximately, introducing only energy error per step.
The Metropolis correction accounts for numerical integration error, ensuring exact invariance with respect to the target.
NUTS removes the need to hand-tune and is the practical choice for most applications.
- Neal, R. M. (2011). MCMC using Hamiltonian dynamics. Handbook of Markov Chain Monte Carlo, 2(11), 2.
- Hoffman, M. D., & Gelman, A. (2014). The No-U-Turn sampler: adaptively setting path lengths in Hamiltonian Monte Carlo. Journal of Machine Learning Research, 15(1), 1593–1623.
- Betancourt, M. (2017). A conceptual introduction to Hamiltonian Monte Carlo. arXiv Preprint arXiv:1701.02434.