CAVI is powerful but requires the variational family to match the conjugate structure of the model. For non-conjugate models — Bayesian logistic regression, neural-network priors, physics-based simulators — no closed-form coordinate updates exist.
Gradient-based variational inference removes the conjugacy requirement by estimating the ELBO gradient with Monte Carlo and optimising with stochastic gradient ascent. This approach goes under several names: black-box VI (BBVI), automatic differentiation VI (ADVI), and fixed-form VI.
The central challenge is differentiating through an expectation:
This chapter covers two gradient estimators that handle this:
Score function estimator (REINFORCE) — broadly applicable but high variance.
Pathwise gradient estimator (reparameterization trick) — lower variance, works whenever we can write with independent of .
We conclude with a worked example: ADVI for Bayesian logistic regression, a non-conjugate model where CAVI has no closed-form updates.
Source
import torch
import torch.distributions as dist
import matplotlib.pyplot as plt
palette = list(plt.cm.tab10.colors)Setup¶
Let denote all latent variables and parameters, and let be the variational family. We maximise the ELBO:
Strategy: assume (unconstrained parameters) and run (stochastic) gradient ascent:
where is an unbiased Monte Carlo estimate of the gradient.
The obstacle: the distribution under the expectation depends on , so we cannot simply move inside the expectation.
The Score Function Estimator (REINFORCE)¶
Derivation¶
Use the log-derivative trick to move the gradient inside the integral:
This gives the score function estimator (a.k.a. REINFORCE Williams, 1992):
Properties:
Works for discrete and continuous — only requires that is differentiable wrt .
Often has high variance in practice.
Variance Reduction: Control Variates¶
Since the score has zero expectation, , we can subtract any baseline without changing the expectation:
Choosing (the running mean of ) can reduce variance substantially.
The Pathwise Gradient Estimator (Reparameterisation Trick)¶
Derivation¶
If we can write where has a distribution independent of (a reparameterisation), then:
The gradient now passes through the expectation cleanly:
Monte Carlo estimate:
In practice: just call .backward() on a Monte Carlo ELBO estimate computed
with reparameterised samples; PyTorch handles the chain rule automatically.
Example: Diagonal Gaussian¶
For :
Non-reparameterisable families: Discrete distributions (Bernoulli, Categorical) cannot be reparameterised in the standard sense. For these, the score function estimator (or straight-through / Gumbel-softmax approximations) must be used.
# ── Variance comparison: score function vs pathwise estimator ─────────────
# Model: observe x=5 from N(theta, 1) with prior N(0, 10^2)
# True posterior: N(mu_post, sigma_post^2) by Bayes rule
# Variational: q(theta; mu, log_sigma) = N(mu, sigma^2)
x_obs = torch.tensor(5.0)
prior_mu, prior_sig = torch.tensor(0.0), torch.tensor(10.0)
lik_sig = torch.tensor(1.0)
# True posterior (Gaussian conjugate)
sigma_post = 1.0 / (1/lik_sig**2 + 1/prior_sig**2).sqrt()
mu_post = sigma_post**2 * (x_obs/lik_sig**2 + prior_mu/prior_sig**2)
print(f'True posterior: N({mu_post.item():.3f}, {sigma_post.item():.3f}²)')
def log_joint(theta):
return (dist.Normal(theta, lik_sig).log_prob(x_obs)
+ dist.Normal(prior_mu, prior_sig).log_prob(theta))
def elbo_and_grads(mu, log_sigma, M, estimator='pathwise'):
sigma = log_sigma.exp()
if estimator == 'pathwise':
# Reparameterise: theta = mu + sigma * eps
eps = torch.randn(M)
theta = mu + sigma * eps
lp = log_joint(theta)
lq = dist.Normal(mu, sigma).log_prob(theta)
elbo = (lp - lq).mean()
elbo.backward()
g_mu, g_ls = mu.grad.clone(), log_sigma.grad.clone()
else: # score function
with torch.no_grad():
theta = dist.Normal(mu, sigma).rsample((M,))
lp = log_joint(theta)
lq = dist.Normal(mu, sigma).log_prob(theta)
h = lp - lq # (M,)
# score wrt mu: (theta - mu) / sigma^2
score_mu = (theta - mu) / sigma**2
# score wrt log_sigma: (theta-mu)^2/sigma^2 - 1
score_ls = (theta - mu)**2 / sigma**2 - 1.0
g_mu = (h * score_mu).mean()
g_ls = (h * score_ls).mean()
elbo = h.mean()
return elbo.detach(), g_mu, g_ls
# Measure gradient variance at the true posterior parameters
torch.manual_seed(42)
mu_val, ls_val = mu_post.item(), sigma_post.log().item()
M_vals = [1, 4, 16, 64, 256]
results = {'pathwise': {}, 'score': {}}
n_trials = 500
for M in M_vals:
for est in ('pathwise', 'score'):
grads = []
for _ in range(n_trials):
mu = torch.tensor(mu_val, requires_grad=True)
log_sig = torch.tensor(ls_val, requires_grad=True)
_, g, _ = elbo_and_grads(mu, log_sig, M, estimator=est)
grads.append(g.item())
results[est][M] = torch.tensor(grads).std().item()
print('\nGradient std (wrt μ) vs. number of MC samples M:')
print(f' {"M":>6} {"Pathwise":>12} {"Score fn":>12}')
for M in M_vals:
print(f' {M:>6} {results["pathwise"][M]:>12.4f} {results["score"][M]:>12.4f}')True posterior: N(4.950, 0.995²)
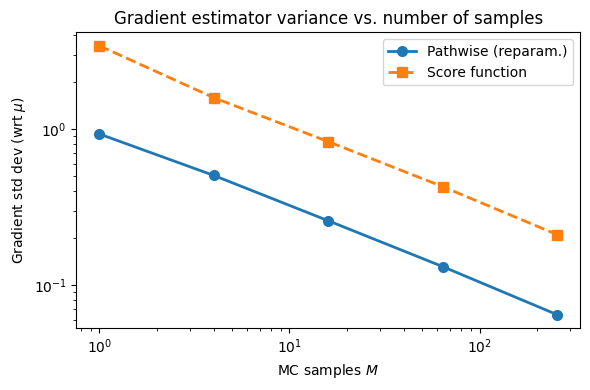
Gradient std (wrt μ) vs. number of MC samples M:
M Pathwise Score fn
1 0.9316 3.4249
4 0.5053 1.5912
16 0.2589 0.8320
64 0.1316 0.4282
256 0.0647 0.2110
Source
fig, ax = plt.subplots(figsize=(6, 4))
ax.plot(M_vals, [results['pathwise'][M] for M in M_vals],
'o-', color=palette[0], lw=2, ms=7, label='Pathwise (reparam.)')
ax.plot(M_vals, [results['score'][M] for M in M_vals],
's--', color=palette[1], lw=2, ms=7, label='Score function')
ax.set_xscale('log'); ax.set_yscale('log')
ax.set_xlabel('MC samples $M$')
ax.set_ylabel('Gradient std dev (wrt $\mu$)')
ax.set_title('Gradient estimator variance vs. number of samples')
ax.legend()
plt.tight_layout()<>:8: SyntaxWarning: invalid escape sequence '\m'
<>:8: SyntaxWarning: invalid escape sequence '\m'
/tmp/ipykernel_2659/3636087577.py:8: SyntaxWarning: invalid escape sequence '\m'
ax.set_ylabel('Gradient std dev (wrt $\mu$)')
ADVI for Bayesian Logistic Regression¶
Model¶
Consider binary observations with -dimensional features . A logistic regression model with Gaussian prior is:
where is the sigmoid. The posterior has no closed form (the likelihood is not conjugate to the Gaussian prior), making CAVI inapplicable.
Variational Family¶
Use a mean-field Gaussian:
where are unconstrained variational parameters ( enforces positivity).
ELBO¶
where . We estimate the expected log-likelihood via a single reparameterised sample () per gradient step, relying on Adam to average out the noise.
torch.manual_seed(305)
# ── Synthetic binary classification data ─────────────────────────────────────
N, D = 200, 4
w_true = torch.tensor([2.0, -1.5, 0.5, -0.8])
X = torch.randn(N, D)
y = dist.Bernoulli(logits=X @ w_true).sample()
print(f'Data: N={N}, D={D}, fraction positive = {y.mean():.2f}')
# ── ADVI with pathwise gradient estimator (Adam) ──────────────────────────────
def advi_logistic(X, y, τ=2.0, M=1, lr=0.05, num_iters=1000, seed=0):
"""ADVI for Bayesian logistic regression.
Variational family: q(w) = N(μ, diag(exp(2ρ)))
Uses the reparameterisation trick + PyTorch autograd.
Parameters
----------
τ : prior standard deviation
M : MC samples per gradient step
"""
torch.manual_seed(seed)
N, D = X.shape
μ = torch.zeros(D, requires_grad=True)
ρ = torch.full((D,), -1.0, requires_grad=True) # log σ, init σ ≈ 0.37
optimiser = torch.optim.Adam([μ, ρ], lr=lr)
elbos = []
for _ in range(num_iters):
optimiser.zero_grad()
σ = ρ.exp() # (D,)
# Reparameterised samples: w = μ + σ * ε, ε ~ N(0,I)
ε = torch.randn(M, D)
w = μ.unsqueeze(0) + σ.unsqueeze(0) * ε # (M, D)
# Expected log-likelihood (averaged over M samples and N data points)
logits = X @ w.T # (N, M)
log_lik = dist.Bernoulli(logits=logits).log_prob(y.unsqueeze(1))
ell = log_lik.sum(0).mean() # scalar
# KL[ N(μ, diag(σ²)) || N(0, τ²I) ] — closed form
kl = dist.kl_divergence(
dist.Normal(μ, σ),
dist.Normal(torch.zeros(D), τ * torch.ones(D))
).sum()
loss = -(ell - kl) # minimise negative ELBO
loss.backward()
optimiser.step()
elbos.append(-loss.item())
return μ.detach(), ρ.exp().detach(), elbos
μ_vi, σ_vi, elbos = advi_logistic(X, y, τ=2.0, M=1, lr=0.05, num_iters=1000)
print('\nADVI posterior means vs. true weights:')
for d in range(D):
print(f' w[{d}]: true={w_true[d]:+.2f}, μ_q={μ_vi[d]:+.4f}, σ_q={σ_vi[d]:.4f}')Data: N=200, D=4, fraction positive = 0.53
ADVI posterior means vs. true weights:
w[0]: true=+2.00, μ_q=+2.2546, σ_q=0.2031
w[1]: true=-1.50, μ_q=-2.0509, σ_q=0.2665
w[2]: true=+0.50, μ_q=+0.6147, σ_q=0.2051
w[3]: true=-0.80, μ_q=-0.5289, σ_q=0.2159
Source
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
# ── Panel 1: ELBO convergence ────────────────────────────────────────────────
axes[0].plot(elbos, lw=1.5, color='steelblue', alpha=0.8)
axes[0].set_xlabel('Iteration')
axes[0].set_ylabel('ELBO')
axes[0].set_title('ADVI: ELBO convergence')
# ── Panel 2: Variational posteriors vs true weights ──────────────────────────
ax = axes[1]
θ_grid = torch.linspace(-4, 4, 300)
for d in range(D):
q_d = dist.Normal(μ_vi[d], σ_vi[d])
ax.plot(θ_grid.numpy(), q_d.log_prob(θ_grid).exp().numpy(),
color=palette[d], lw=2, label=f'$q(w_{d+1})$')
ax.axvline(w_true[d].item(), color=palette[d], lw=1.5, ls='--', alpha=0.7)
ax.set_xlabel(r'$w$')
ax.set_ylabel('Density')
ax.set_title('Variational posteriors $q(w_d)$\n(dashed = true values)')
ax.legend(fontsize=8)
# ── Panel 3: Predictive accuracy ─────────────────────────────────────────────
ax = axes[2]
# Predict using posterior mean
logits_train = X @ μ_vi
y_pred = (logits_train > 0).float()
acc_vi = (y_pred == y).float().mean().item()
# Compare to MLE / MAP
w_mle = torch.linalg.lstsq(X, y.float()).solution
y_pred_mle = ((X @ w_mle) > 0.5).float()
acc_mle = (y_pred_mle == y).float().mean().item()
# MC predictive
M_pred = 1000
ε_pred = torch.randn(M_pred, D)
w_samp = μ_vi.unsqueeze(0) + σ_vi.unsqueeze(0) * ε_pred # (M_pred, D)
prob_pred = dist.Bernoulli(logits=X @ w_samp.T).probs.mean(1).numpy()
ax.hist(prob_pred[y == 1], bins=20, alpha=0.6, color=palette[0], label='True y=1')
ax.hist(prob_pred[y == 0], bins=20, alpha=0.6, color=palette[1], label='True y=0')
ax.set_xlabel('MC predictive probability $p(y=1 | x_n)$')
ax.set_ylabel('Count')
ax.set_title(f'MC predictive probabilities\n(train accuracy: {acc_vi:.2f})')
ax.legend(fontsize=9)
plt.tight_layout()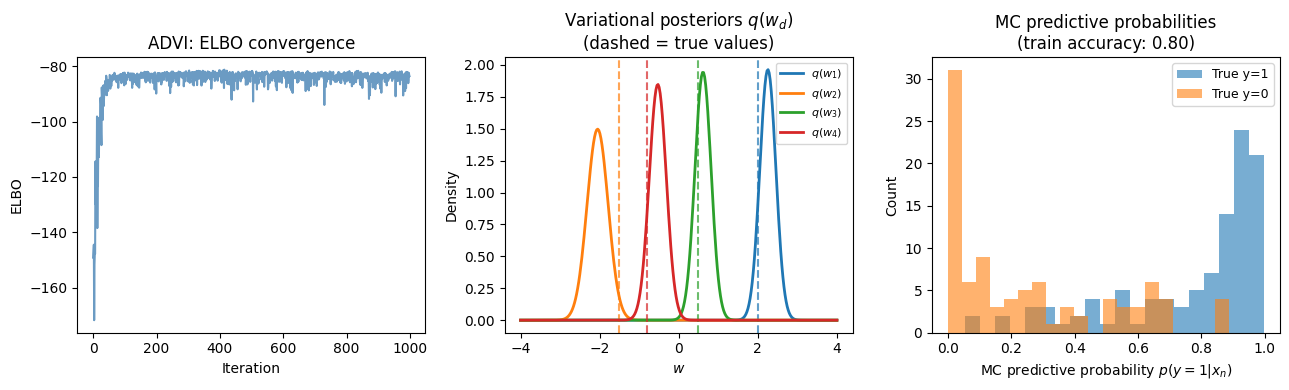
Mini-Batches and Stochastic Optimisers¶
Mini-Batch ELBO¶
When is large, each gradient step over the full dataset is expensive. We can subsample a mini-batch and scale up the log-likelihood:
giving an unbiased estimate of the ELBO and its gradient.
Stochastic Optimisers¶
Basic SGD often converges slowly. Several adaptive-step methods work well for ELBO optimisation:
| Method | Update rule | Notes |
|---|---|---|
| SGD Robbins & Siegmund, 1971 | Simple; needs careful schedule | |
| AdaGrad Duchi et al., 2011 | Per-parameter | Good for sparse gradients |
| Adam Kingma & Ba, 2014 | Momentum + adaptive scale | Default choice; robust |
Robbins-Monro conditions for SGD convergence: the step sizes must satisfy and .
Adam is the practical default for ADVI — it rarely requires step-size tuning and handles the varying scales of and automatically.
Conclusion¶
Gradient-based VI removes the conjugacy requirement of CAVI, enabling variational inference in any model where the log joint is differentiable.
| Score function | Pathwise (reparam.) | |
|---|---|---|
| Requires | Differentiable wrt | Differentiable reparameterisation |
| Works for discrete ? | Yes | No (without approximation) |
| Variance | High | Low |
| Implementation | Manual score computation | Automatic (.backward()) |
Key takeaways:
The log-derivative trick converts into an expectation computable by MC — but with high variance.
The reparameterisation trick decouples sampling noise from , moving the gradient inside the expectation and drastically reducing variance.
In PyTorch, reparameterised VI is nearly automatic: construct the ELBO using
.rsample(), call.backward(), step an optimiser.Adam is the practical default; a single MC sample per step () usually suffices when paired with an adaptive optimiser.
The KL term between two Gaussians (or other conjugate pairs) can be computed analytically, reducing MC noise further.
- Williams, R. J. (1992). Simple statistical gradient-following algorithms for connectionist reinforcement learning. Machine Learning, 8(3), 229–256.
- Robbins, H., & Siegmund, D. (1971). A convergence theorem for non negative almost supermartingales and some applications. In Optimizing methods in statistics (pp. 233–257). Elsevier.
- Duchi, J., Hazan, E., & Singer, Y. (2011). Adaptive subgradient methods for online learning and stochastic optimization. Journal of Machine Learning Research, 12(7).
- Kingma, D. P., & Ba, J. (2014). Adam: A method for stochastic optimization. arXiv Preprint arXiv:1412.6980.
- Blei, D. M., Kucukelbir, A., & McAuliffe, J. D. (2017). Variational Inference: A Review for Statisticians. Journal of the American Statistical Association, 112(518), 859–877.
- Mohamed, S., Rosca, M., Figurnov, M., & Mnih, A. (2020). Monte Carlo Gradient Estimation in Machine Learning. Journal of Machine Learning Research, 21(132), 1–62.